Making a run for it: escaped farmed Atlantic salmon integrating with wild populations
By Robbie Roemer, SRC master’s student
Atlantic salmon (Salmo salar) as their name implies, are primarily found in northern Atlantic waters and are classified as androminous (living in the sea, and returning to freshwater to spawn). Known to be a popular recreational sport fish, this largest species found in the genus Salmo is prized for its table fare and thus, faces heavy commercial fishing pressure. This species is particularly sensitive to habitat alteration and human influence (Staurnes et al. 1995; Kroglund et al. 2007) and coupled with the high commercial demand, has seen significant historical declines over the last half century. These declines have led to substantial increases in aquaculture farming techniques where salmon are raised in pens on the very same waters utilized by native, wild populations to spawn. Breeding and farming programs have greatly altered the genetic makeup of Atlantic salmon as commercial enterprises target specific characteristics such as: larger total size, faster growth rates, efficient food utilization, and meat quality. But what happens to the inevitable large quantity of farm “escapees”?
![Atlantic salmon are popular sport fish in Norway and beyond [Image by Vetle Kjærstad]](https://sharkresearch.rsmas.miami.edu/wp-content/uploads/2017/04/Roemer_Figure_1-300x225.jpg)
Atlantic salmon are popular sport fish in Norway and beyond [Image by Vetle Kjærstad]
So why is the genetic introgression or “mixing “of farmed salmon to wild salmon ecologically important? The main concerns by the authors regarding genetics are the loss of genetic variation within a population, the loss of genetic variation between populations, and the loss of overall animal ecological fitness. It has long been shown that farmed salmon have much lower genetic variation compared to their wild counterparts ((Mjølnerød et al. 1997; Skaala et al. 2004, 2005; Karlsson et al. 2010). In addition, substantial loss of ecological fitness has been documented in farm-raised salmon. If wild to farmed genetic introgression continues at this rate, it is feared wild salmon populations will too lose genetic attributes, all of which are critical in sustaining healthy, disease-free, wild salmon populations.
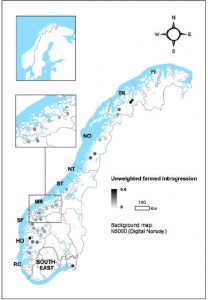
Map of Norway showing rivers with farmed genetic introgression (Karlsson et al. 2016).
This research has real-world applications, as many hydropower companies that alter the natural state of rivers, and reduce natural productivity of native salmon compensate this “offset” by releasing farm raised fish into the river system. In the western United States, native cutthroat trout are facing a similar threat, as genetic introgression with rainbow trout is occurring at a rapid rate. It has been proposed to list the few remaining genetically “pure” populations of cutthroat trout under the Endangered Species Act (ESA). Similar proposals have been made to Atlantic salmon, even going so far as to list farm-raised salmon a different species, and treating farm raised “escapees” as an exotic species, to help deter genetic hybridization and introgression with wild populations.
One positive finding within the study was the lowest genetic introgression rates were located within Norwegian nationally protected lands (National Salmon Rivers and National Salmon Fjords), thereby demonstrating the ecological importance of preserved lands to wildlife populations. Indeed, there is no clear, sound solution to this problem, especially as the numbers of salmon farms are increasing globally. However, it is clear that at the present time, near-zero limits are the only viable solution to protect the genetic integrity of wild Atlantic salmon populations.
Works Cited
Karlsson, S., Diserud, O.H., Fiske, P. and Hindar, K., 2016. Widespread genetic introgression of escaped farmed Atlantic salmon in wild salmon populations. ICES Journal of Marine Science: Journal du Conseil, 73(10), pp.2488-2498.
Kroglund, F., Rosseland, B.O., Teien, H.C., Salbu, B., Kristensen, T. and Finstad, B., 2007. Water quality limits for Atlantic salmon (Salmo salar L.) exposed to short term reductions in pH and increased aluminum simulating episodes. Hydrology and Earth System Sciences Discussions, 4(5), pp.3317-3355.
Staurnes, M., Kroglund, F. and Rosseland, B.O., 1995. Water quality requirement of Atlantic salmon (Salmo salar) in water undergoing acidification or liming in Norway. Water, Air, & Soil Pollution, 85(2), pp.347-352.


